Looking at the Futures - June 04, 2020
Reviewing positions...
Last Friday was the last trading day of the May BTC derivative contracts on the CME. So now is a good time to look back at the action and see how traders are positioned for the months to come.
Looking at the Futures is a weekly newsletter where we are making sense of the Bitcoin derivatives market. Emphasis is put on the CME products as they dominate the field for institutional investors. But we’ll occasionally look at other venues too.
Subscribe now and don’t miss anything!
These days it’s all about $10,000. For over a month now Bitcoin is stuck trading in a range. Roughly between $8,500 and $10,000 with some brief excursions slightly above or below that range. With such a long time spent right under the psychological $10,000 mark you can guess that something is brewing.
And that something is bulls sweat!

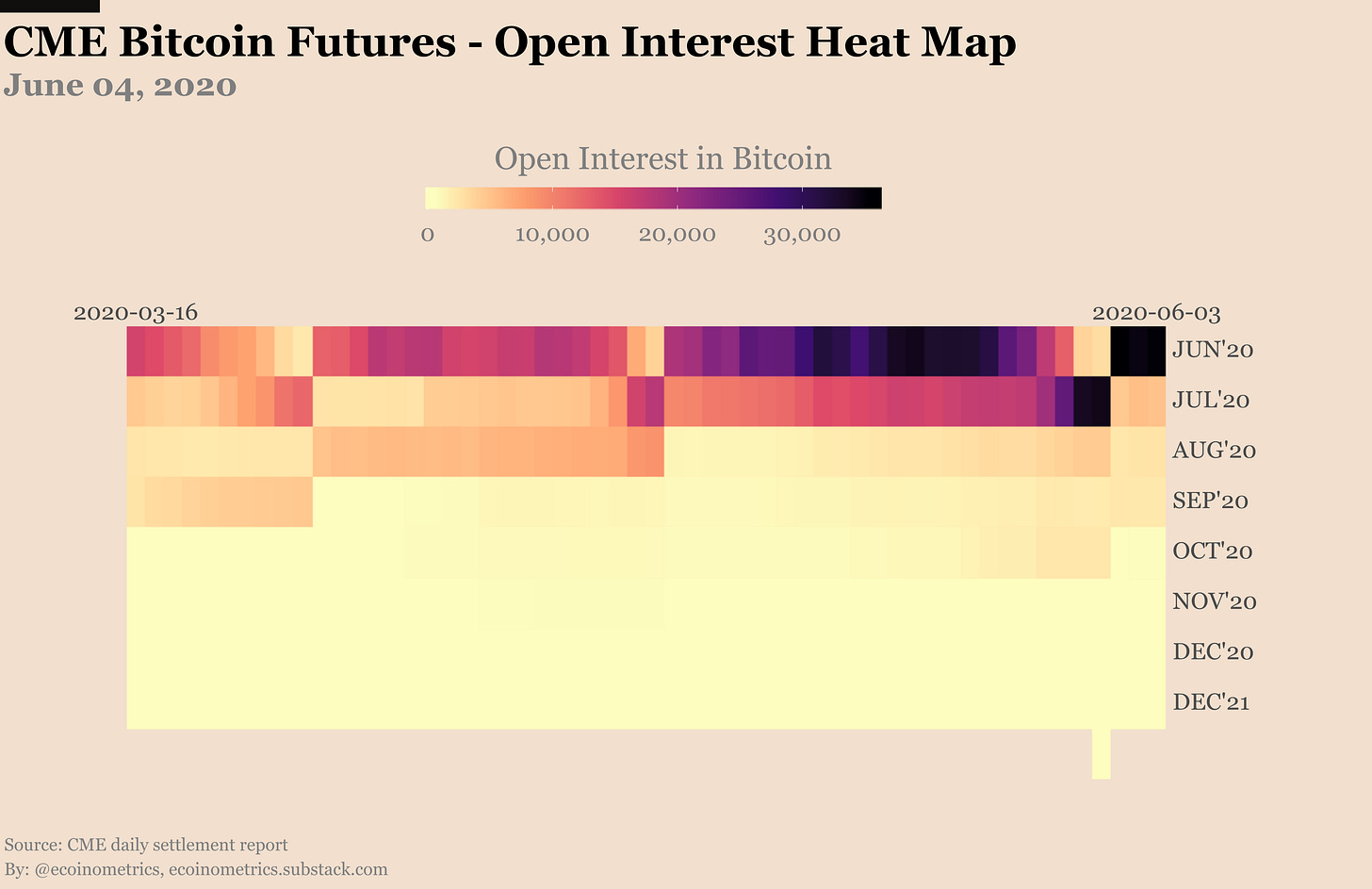
Look at the futures open positions in conjunction with the price action. We’ve had a big increase in open interest and very little liquidations even when the price looked like it was going to dip lower. Even a failed break out above $10,000 this week didn’t result in many people giving up on the dream of the bull market.

Actually when you look at the heat map of open interest broken down by contract month it is clear that we the number of positions on the front month is already back to the record high levels of last month. At the time I’m writing this letter there is about 35,000 Bitcoin worth of contracts on Jun’20.
To me it feels like traders are still waiting for Bitcoin to push significantly above $10,000. If you think about it in terms of a momentum trade that probably makes sense.
From the bottom of March to now Bitcoin is up +100%. It is still trading above its 200 days moving average. So if you are in it for the trend then there is no point going out now. That might change if Bitcoin fails to break $10,000 for a significant amount of time though.
Add to that the fact that $10,000 as a round number is a psychologically attractive resistance level to break and you most likely have a full picture of the current sentiment in the market.

Things are even more obvious when you look at the positioning on the options market after the May contract expired. First of all the market continues to be totally driven by the activity on the calls.
Only a few days after the May contract expired we are almost back to all time high open interest on the options market. About 25,000 Bitcoin worth of call contracts open in total.

Most of those calls are between the $10,000 to $15,000 strikes. But not all of them are positioned on the June contract. Actually we can have a slightly clearer picture if we decompose the positions distribution between June and July where the bulk of the contracts reside.
Take a look.



I’d interpret that as a number of investors making some reasonable bets on where Bitcoin could go in the next couple of months. As I write those lines Bitcoin stands at $9,500. So we have something like:
$10,500 is a +10% move.
$11,000 is a +16% move.
$11,500 is a +21% move.
$12,000 is a +26% move.
$13,000 is a +37% move.
Considering the typical volatility of Bitcoin and the speed at which it recovered 100% from it low of March, betting on a +10% to +37% move in the next two months is totally reasonable.
That’s why we have seen bull calls popping up since Bitcoin closed in on $10,000. If you count on a measured move you’d buy some bull call spreads to capture as much profit as possible.
For those of you who aren’t familiar with what a bull call spread is for check out the explanation below for a brief description.

This is different from what we have seen in the past. Remember only a few months ago it was more common to see traders buying far out of the money calls in anticipation of some wild ride after the halving. But in terms of trading activity that was child’s play. The addition of a more conservative approach to strategic positioning is what brought the big trading volumes to the CME options market.
So here it is for the state of how traders are positioned. But what are the traders doing category by category? As usual the CFTC commitment of traders data is lagging by one week but there is always useful information to glean from it.
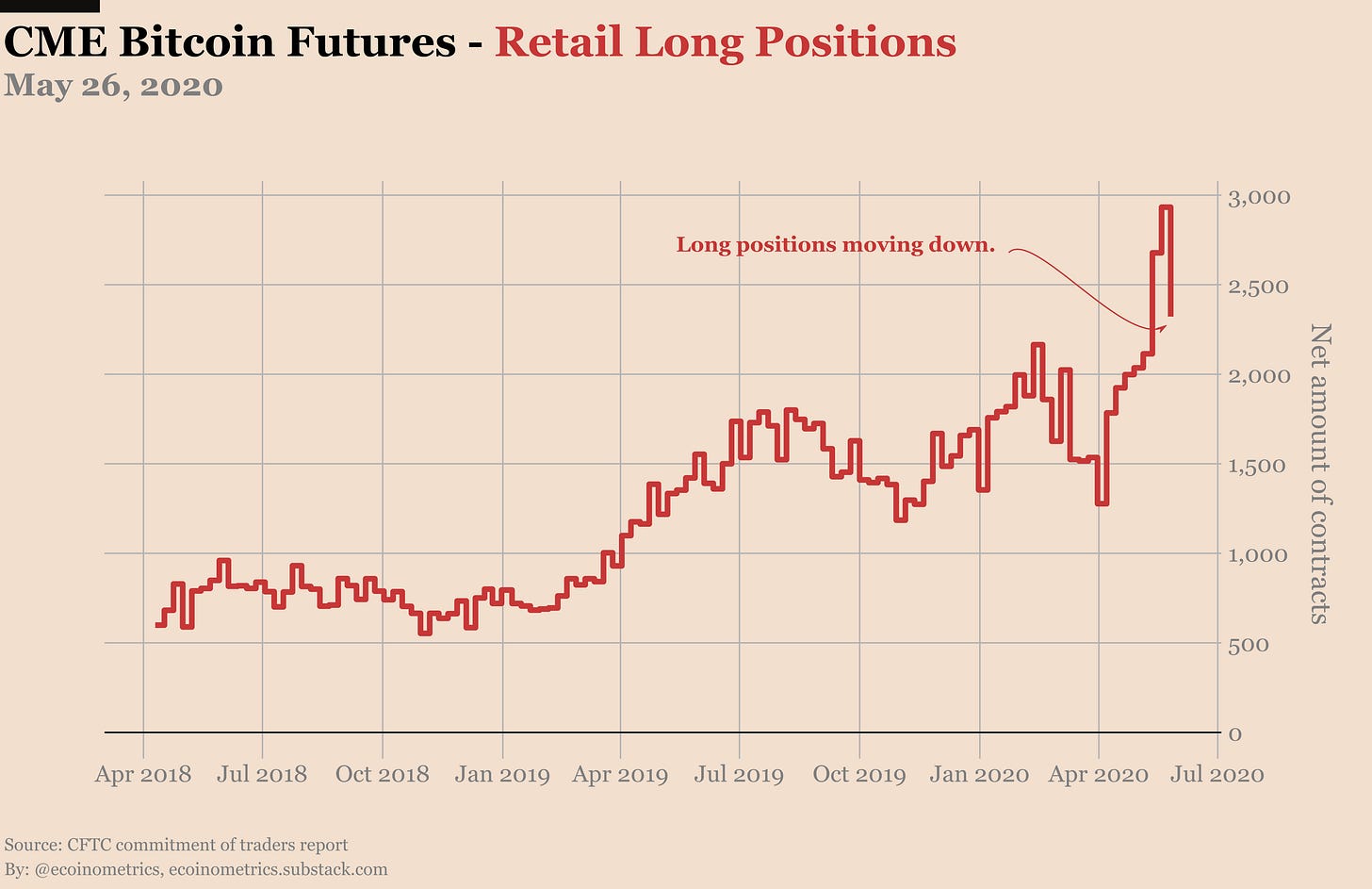
First of all the retail traders. Obviously they are still net long Bitcoin. However they aren’t nearly as long as they were a couple of weeks ago. We are now all the way down to the previous highs of January.
And is it that they suddenly came in with new shorts or is it simply the result of liquidation of the longs? Well it is both! There was about 500 long contracts liquidated and 300 shorts added.
Some retail traders don’t expect $10,000 to be broken so soon apparently.


When it comes to the smart money the situation is reversed. Both long and short positions have been liquidated. However more shorts than longs were closed. Overall the smart money is still short Bitcoin though.

That’s it for this week. In the long term Bitcoin is moving higher. But how long that will take might significantly change how traders are positioned on the derivatives markets. For now the general sentiment is bullish. But get a few more false breaks above $10,000 and that could turn sour.
Let’s see how it plays out.
If you liked this article please share and subscribe to help the Ecoinometrics newsletter grow!

